The natural pyrazolotriazine pseudoiodinine from Pseudomonas
发布日期:2023-03-19 20:39 浏览次数:
Ruihuan Yang, Qing Shi, Tingting Huang, Yichao Yan, Shengzhang Li, Yuan Fang, Ying Li, Linlin Liu, Longyu Liu, Xiaozheng Wang, Yongzheng Peng, Jiangbo Fan, Lifang Zou, Shuangjun Lin & Gongyou Chen
Nature Communications
Natural products largely produced by Pseudomonads-like soil-dwelling microorganisms are a consistent source of antimicrobial metabolites and pesticides. Herein we report the isolation of Pseudomonas mosselii strain 923 from rice rhizosphere soils of paddy fields, which specifically inhibit the growth of plant bacterial pathogens Xanthomonas species and the fungal pathogen Magnaporthe oryzae. The antimicrobial compound is purified and identified as pseudoiodinine using high-resolution mass spectra, nuclear magnetic resonance and single-crystal X-ray diffraction. Genome-wide random mutagenesis, transcriptome analysis and biochemical assays define the pseudoiodinine biosynthetic cluster as psdABCDEFG. Pseudoiodinine biosynthesis is proposed to initiate from guanosine triphosphate and 1,6-didesmethyltoxoflavin is a biosynthetic intermediate. Transposon mutagenesis indicate that GacA is the global regulator. Furthermore, two noncoding small RNAs, rsmY and rsmZ, positively regulate pseudoiodinine transcription, and the carbon storage regulators CsrA2 and CsrA3, which negatively regulate the expression of psdA. A 22.4-fold increase in pseudoiodinine production is achieved by optimizing the media used for fermentation, overexpressing the biosynthetic operon, and removing the CsrA binding sites. Both of the strain 923 and purified pseudoiodinine in planta inhibit the pathogens without affecting the rice host, suggesting that pseudoiodinine can be used to control plant diseases.
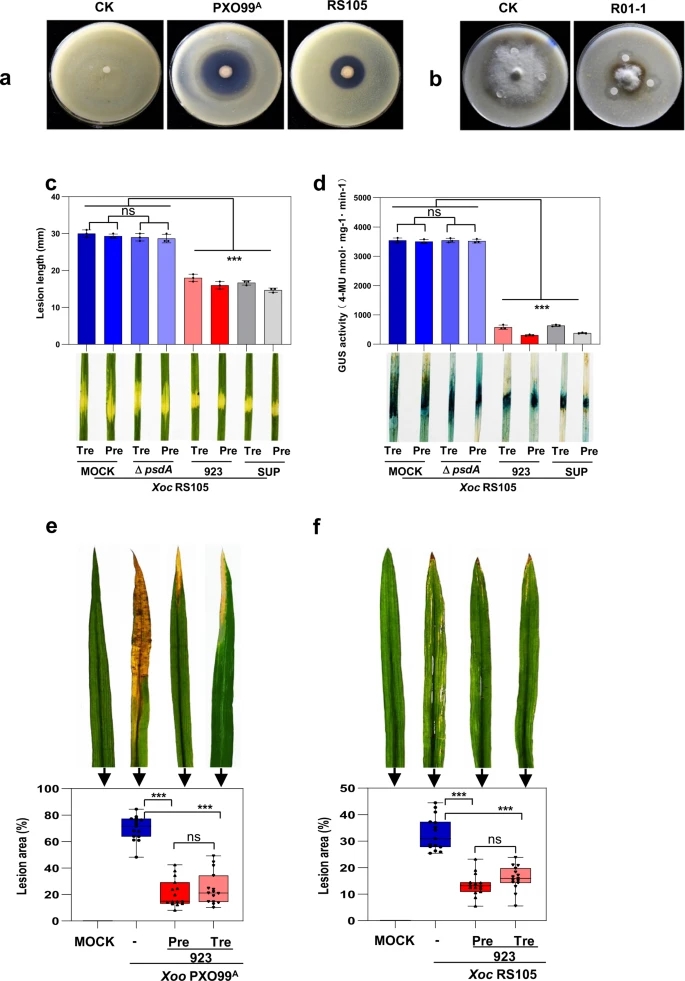 Fig. 1: Antagonistic and biocontrol activity of P. mosselii 923.
Fig. 1: Antagonistic and biocontrol activity of P. mosselii 923.
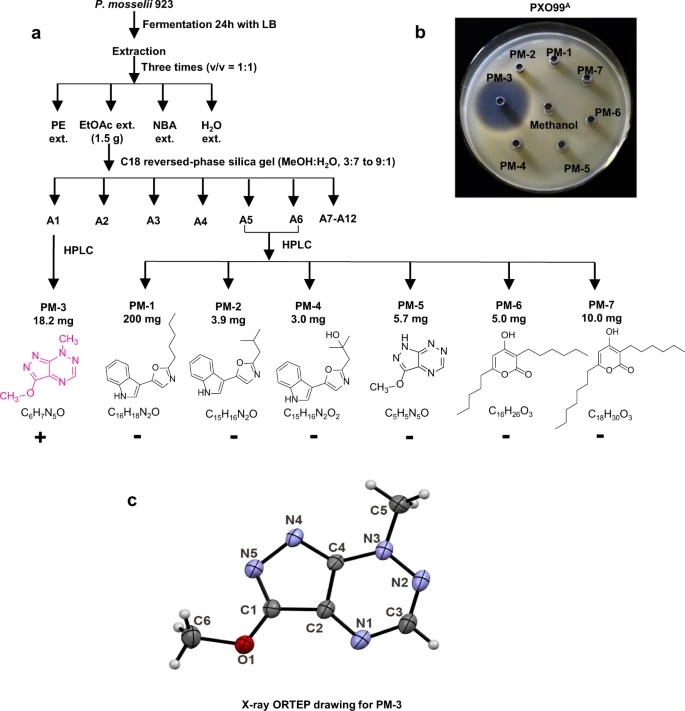
Fig. 2: Extraction and purification of PM-3 (pseudoiodinine).
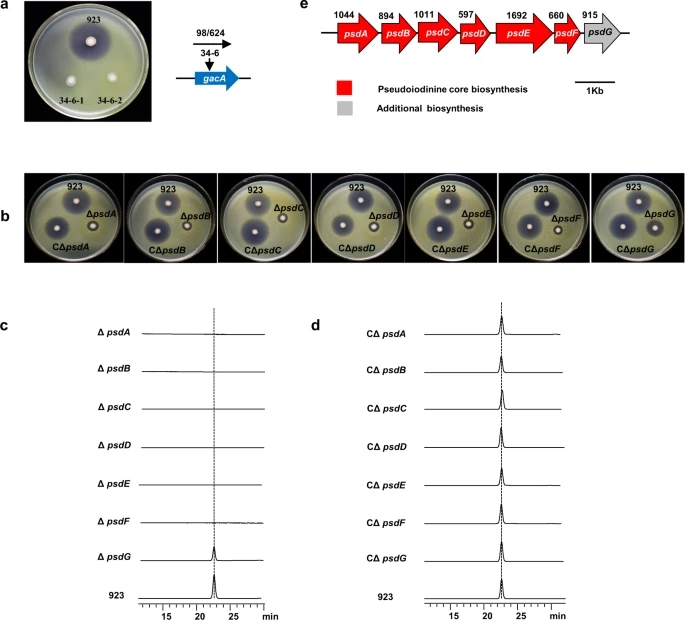
Fig. 3: Genetic determinants of pseudoiodinine biosynthesis.


